Without a single test for ALS, the diagnosis of the disease is by exclusion. Clinicians monitor people with ALS instead using questionnaires and rudimentary clinical tools. Few treatment options are available on pharmacy shelves.
Emerging technologies, however, promise to help change that. Cutting-edge imaging tools are beginning to reveal the neuronal circuitry destroyed by ALS – paving the way toward identifying and tracking the disease. The advent of next-generation sequencing technologies has led to an explosion of ALS genes - sparking new ideas for treatment strategies that target emerging disease mechanisms. And, mind-melding brain machine interfaces hope to help people keep moving or to walk again.
Clinicians and scientists gathered at the 2013 Meeting of the American Association for the Advancement of Science (AAAS) in Boston to discuss the latest technologies and the challenges to bring them into general clinical practice.
Of Genes and Genomes
ALS is a complex heterogeneous disease. Many people experience their first signs of ALS in their 50s and survive 2 – 5 years. But others get ALS earlier and/or live longer with the disease.
The reason, suspect scientists, lies in part in their genes. Certain genetic differences called modifiers influence the time of onset and duration of ALS. A few of these “variants” have been uncovered. But many more are still to be discovered.

Decoding ALS? Researchers are working hard to sequence the genomes of people with ALS in hopes to develop better tools to identify and treat them. Image: Roy Kaltschmidt, Lawrence Berkeley National Laboratory.
A large team of US researchers led by Hudson Alpha’s Rick Myers PhD is now hard at work sequencing the genomes of 1000 people with ALS. The genetic differences detected might help scientists identify new targets and therapies for the disease.
The project is one of a number of ongoing efforts around the globe that hope to create better tests and better treatments for people with a wide-range of medical conditions. But how this genetic information will translate to better diagnosis and management of disease remains hotly debated according to physicians at AAAS13.
A key challenge according to University of Pennsylvania School of Medicine’s Reed Pyeritz MD PhD is the inherent uncertainty of today’s genomic medicine. An estimated 4 million variants can be found within our genomes. Many of these genetic differences have never been seen before. And, most of these changes are of uncertain significance. Clinicians simply do not know what most of these changes mean in terms of health and disease according to University of North Carolina School of Medicine’s James Evans MD PhD.
“I think interpreting variants is the single biggest challenge in the next decade,” says Evans.
Unlike an X-ray that indicates a broken bone or a critical infection, clinicians are unable to fully decipher the information hidden in our genomes. More than 1500 variants have been linked to increased risk of developing 200 complex genetic diseases. Clinicians are simply unsure which of these changes signal that their patients are reaching the danger zone – making their interpretation more of a “parlor game” says Evans.
To sequence or not to sequence? Video: Mount Sinai School of Medicine.
“Our ability to dissect the clinical genome isn’t good yet,” says Evans. “We simply do not know how to use [genomics] in medicine at this point.”
But while some clinicians demand more evidence before implementing these tools, others are forging ahead developing guidelines to use them. The reason: whole genomic analysis is already available in the clinic and is soon to become general practice.
“Even though the data is insufficient, clinicians must still provide advice, patients must still make choices and policy makers must still make policies,” says Harvard Medical School’s Robert Green MD MPH, director of the Genomes2People project, quoting from a 2009 report by the US Prevention Task Force.
The Harvard Medical School team is developing a one page “General Genome Report” that includes potentially key genetic changes and pharmacokinetic status to help inform drug recommendations.
The report is one of growing number that aims to help doctors provide better care for their patients. In June 2012, Foundation Medicine introduced a next-generation sequencing-based tumor test that seeks out the usual suspects - mutations in 200 oncogenes - in people with cancer. The results, relayed in a short report, hopes to help oncologists decide on the most appropriate treatment strategies for their patients – including those currently being tested in clinical trials.
Meanwhile, Harvard University’s George Church PhD introduced a hospital-friendly supercomputer (Knome's Knosys 100) in September 2012 that aims to enable clinicians to zero in on potentially key variants likely linked to their patients' disease.
Man and machine
Elsewhere across the globe, a growing number of engineers and material scientists are developing devices that tap into the nervous system in hopes to provide better care for people with neurological conditions including ALS.
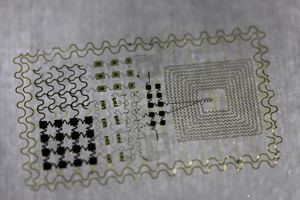

Tattoo Nation? Researchers are developing small, flexible, skin sensors to measure electrical activity of the brain and the muscles. Image: John Rogers PhD, University of Illinois at Urbana-Champaign.
A regular doctor’s visit might soon be a lot more comfortable thanks to electronic tattoos being developed by University of California San Diego bioengineer Todd Coleman PhD and University of Illinois Urbana-Champaign material scientist John Rogers PhD. The peel and stick sensors, an alternative to itchy electrodes and uncomfortable needles, works much like an EEG ECG or an EMG – monitoring the electrical impulses of the brain, heart or muscles. Simply apply when wet much like temporary tattoos. Electronic tattoos are currently being explored for a number of uses in the clinic according to Coleman including tracking muscle function in people with ALS.
A full body suit that aims to turn thoughts into actions being developed by an international team led by Duke University neuroengineer Miguel Nicolelis MD PhD hopes to enable paralyzed people to walk again. The brain machine interface-based device called an exoskeleton works by recording impulses from thousands of neurons in multiple regions of the brain. “Plasticity takes care of the rest,” says Nicolelis.
The robotic suit is expected to be unveiled at the 2014 World Cup in Brazil. A paraplegic, wearing the suit, will have one of the first shots on goal during the opening game. A prototype is currently being tested in monkeys. “It is not brain-controlled yet but we are getting there,” says Nicolelis.
***
Making Connections
ALS is a progressive neurodegenerative disease that leads to muscle weakness, paralysis and ultimately respiratory failure. But where ALS starts and how the disease spreads remains an open question.
A growing number of studies suggest that ALS is a systems failure – a series of neural networks go offline leading to a loss of muscle function. Researchers are hard to work to identify and map these networks in people with ALS in hopes to uncover the “hubs” of their disease. The results might help clinicians identify people with ALS earlier and track their progression.
“If we know where the disease begins, we can predict where the disease will go,” explains University of California San Francisco (UCSF) neurologist Bill Seeley MD.
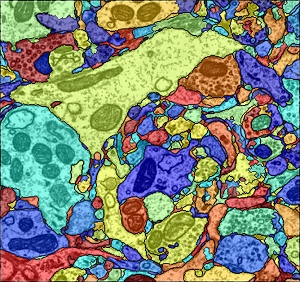
Tracing the cause? Researchers can now identify individual cells including neurons, astrocytes and microglia in slices of brain tissue enabling them to capture snapshots of the brain at synapse resolution. Image: Amelio Vázquez-Reina PhD, Human Connectome Project.
The strategy: Image people with ALS by resting-state functional MRI. Identify the troublespots (networks affected) by comparing them to healthy people. Locate the "epicenters" using network trackbacks.
The approach is now helping to unravel a number of neurodegenerative diseases including Alzheimer’s disease and frontotemporal dementia (FTD).
But the circuitry ultimately identified by these methods is only roughly mapped out in the brain – the locations instead deduced by areas of reduced or increased brain activity. To truly snuff out the exact source of their disease, scientists need a detailed wiring diagram of the brain: a map of the connectome.
Researchers are already beginning to do just that. The Human Connectome Project (HCP), led by scientists at Massachusetts General Hospital and the University of California Los Angeles, aims to map the brain’s superhighways in 1,200 people.
The results are to be made available online at the HCP website. Quarterly releases are expected starting in May 2013.
“We hope that this will lead to a better understanding of brain circuitry in health and disease,” says HCP investigator and neuroscientist Steve Petersen PhD of the Washington University School of Medicine.
The initiative, kickstarted in 2011, aims to capture the brain using cutting edge imaging techniques at multiple levels - including the creation of a wiring diagram detailing neuron-neuron connections.
The electron microscopy-based technique, developed by a team led by Harvard University's Jeff Lichtman MD PhD, operates much like a movie projector in reverse – reconstructing regions of the human brain at synapse resolution. The resulting wiring diagrams include key targets of neurodegenerative disease including neurons, mitochondria, glia and synaptic vesicles.
The goal is to identify key circuits damaged in neuropsychiatric diseases including autism and schizophrenia to get a better idea how to treat them. But this technique is expected to help scientists unravel many neurodegenerative diseases – including ALS.
“Neurological diseases look like something at this level,” says Lichtman.
Clinical Trial and Error
Researchers are unraveling ALS at increasing speed. New medicines are being developed to target these emerging disease mechanisms. And, existing therapies are being repurposed to bring medicines more quickly to the clinic.
But the field continues to be plagued with disappointments. Ceftriaxone and dexpramipexole, posting more than a 30% drop in disease progression at phase II, failed at phase III. Treatments for people with ALS continue to be limited. Mitsubishi Tanabe Pharma’s Radicut (edavarone) is one of the only drugs currently being tested at phase III around the globe.
People with ALS are not alone. ALS is one of hundreds of diseases without an effective treatment or cure. And, more than 90% of drugs for these diseases fail in clinical trials according to Johns Hopkins University School of Medicine toxicologist Thomas Hartung MD PhD. “We are putting are money on the wrong horses.”

Next top ALS model? Researchers are working hard to develop mouse models of ALS that resemble more common forms of the disease in hopes to identify more effective medicines. Image: Wellcome Library, London.
A key problem according to Hartung is how emerging medicines are developed at the preclinical stage. Most drugs cannot be independently validated according to a growing number of studies. And, others are found to be intolerable or unsafe. The reason says Hartung is the lack of sufficiently rigorous safety and efficacy testing practices using validated methods.
“We have to praise our animal models to get them published,” says Hartung. “[But] we need to understand that they have limitations.”
Choosing the right system and the right methods to push forward drugs into the clinic, however, is not the only obstacle according to Anne Glover CBE FRSE FAAM, Chief Scientific Advisor of the European Union’s European Commission. There is considerable red tape. And, results from completed clinical trials are not always shared. “I want to see the data,” says Glover.
This transparency according to National Institutes of Health’s Wilson Compton MD MPE is essential to allow independent analysis of clinical trial results. An analysis that is needed to ensure the safest and the most promising medicines are pushed forward into the clinic as quickly as possible. And, according to Glover, is needed to keep the faith in the drug approval process.
“We need clinical trials. We need them to be the best that they can be,” says Glover.
The move is gaining momentum throughout the globe. In the US, Congressman Ed Markey introduced a bill in August 2012 called the Trial and Experimental Studies Transparency (TEST) Act which mandates that the registration of all clinical trials on the ClinicalTrials.gov website within one month after being funded by NIH and the posting of results within 1 year of completion. A bill endorsed by the New England Journal of Medicine. In Europe, British MEP Glenis Willmott is pushing hard in the European Parliament for legislation that requires clinical trial sponsors to file a “Clinical Trials Report” that contains study results or face fines. And, last month, the industry group Association of Biotech Led Enterprises (ABLE) in Mumbai pledged to help make results available and accessible from certain clinical trials taking place in India.
”Nothing is risk free,” says Danish Ministry of Science, Technology and Innovation’s Klaus Block PhD. “Openness and trust is absolutely essential to improve outcomes of clinical trials.”